k-means clustering

k-means clustering is a neat way to find ‘centers of density’ in a group of points. It’s useful for applications like finding natural groups, simplifying visualizations, and summarizing data.
There are plenty of libraries that implement the algorithm in Javascript, R and more. While I’ve been working on an implementation with some interesting properties, this article is about understanding the algorithm.
Since it’s tricky to visualize the inner workings of the 2D form, let’s look at k-means in one dimension.
What are cluster means?
A cluster mean is a new point that represents the center of a cluster in the data. k-means aims to equally distribute the cluster means by minimizing the aggregate distance between means and values - technically the within-cluster sum of squares.
Sum of squares should be familiar from simple linear regression, which also tries to minimize an aggregate distance, but between points and a line rather than points and other points.
This algorithm works by refining its guesses for cluster means iteratively until they’re ‘good enough’ - they minimize the sum of squares.
The Algorithm
The most popular algorithm for calculating k-means is Lloyd’s algorithm - a heuristic: it comes up with a viable solution but not necessarily the best one. Calculating the true, perfect k-means would take a whole lot longer, and offer only a marginal improvement. It’s also iterative: it refines a guess repeatedly until it’s ‘good enough.’
Input

cluster_means = kmeans(dataset, k)The input is a d-dimensional dataset of length n, and a k value - which indicates how many cluster means should be produced.
Selecting the initial means
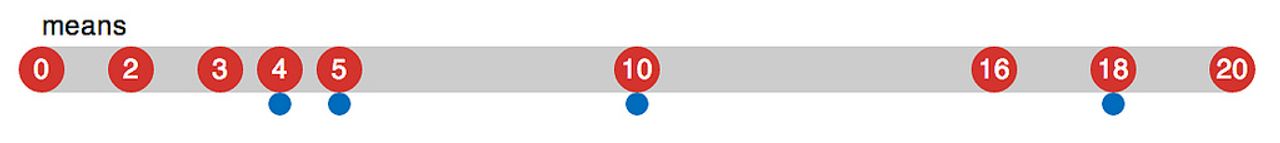
The algorithm iteratively improves the cluster means - but where do those come from? They’re chosen randomly from the data - for k desired means, we choose k values from the n values in the data.1
These values are the potential means - hence k-means. But since we just chose them randomly, they likely aren’t that good - they might even be terribly lopsided, inhabiting only one small part of the dataset. This doesn’t matter - doing the next two steps repeatedly will make them drift towards much better values.
Assigning points to means
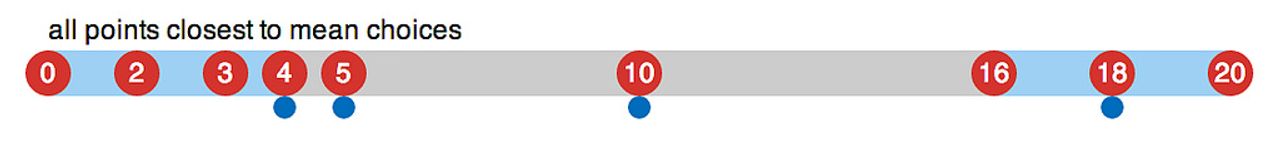
Next, each point is grouped with the closest mean - effectively creating a voronoi tessellation with them as centers.
Comparing every point to every center can be very slow. Thus some implementations use shortcuts, like the polymaps implementation uses a k-d tree to speed things up.
The distance between points is measured as the euclidean distance. This is kind of neat in that it can work in many dimensions. My kmeans implementation has the following distance function that handles arbitrary dimensions.
function dist(a, b) {
var d = 0;
for (var i = 0; i < a.length; i++) {
d += Math.pow(a[i] - b[i], 2);
}
return Math.sqrt(d);
}At the end of this step, every point in your data is in a group belonging to a potential mean.
Deriving new means
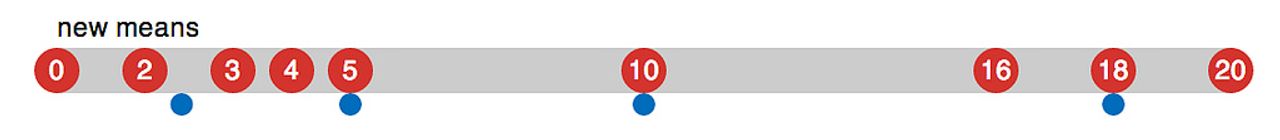
This one’s simple - just take the centroid of each group of points. In one dimension, this is just the average of all of the points. In two, it’s the average of each dimension.
The resulting averages are the new cluster means, and you can now go back to Assigning points to means and repeat the process until the cluster mean values stabilize.
Boundary Cases
If k is equal to n (the size of the data itself), the output is the same as the input, since the distance between each point and the representative mean is zero.
If k is one, then the eventual output will be the global mean - the average of all the values.
How it feels
Play with k-means: this shows the calculation, step by step, of cluster means from input values. Drag numbers in the first line to change the input. This requires a modern browser.
Applications of kmeans
Wikipedia has a longer but less specific list
In cartography kmeans can be used for finding centers of density in 2D space. However, it’s a different problem and a different solution than the more common point ‘decluttering’ that intends to reduce visual noise and overlapping shapes: k-means is not sensitive to symbolization, so cluster points can still overlap.
The algorithm can be used to select palettes for color reduction in images. What’s cool about that application is that the RGB color spaces is a three-dimensional cube, and thus the clustering happens in three dimensions.
See Also
- This actually turned out to be a bit of a stumper - the algorithm in the polymaps implementation is suboptimal for when your k value approaches n, and I didn’t want to settle for a random-sorting and slicing approach. I ended up implementing a Javascript version of Floyd’s algorithm for random subsets.